| Video Discription |
DNA replication. My website, science with susanna.com has the blank drawing to accompany this video as well as practice materials to quiz yourself.
DNA replication produces two identical molecules of DNA from the original. This process occurs during S phase of interphase.
1 tightly packaged chromosome contains 50 - 250 million base pairs. Here’s a piece of it stretched out for us so that we can study this process of DNA replication. The DNA code consists of matching nucleotides - adenine with thymine; and guanine with cytosine.
Here’s one of these millions of base pairs. Base pairs are bonded together with hydrogen bonds.
DNA is oriented with anti-parallel strands. One end is oriented 5 prime to 3 prime, and the complementary strand is oriented 3 prime to 5 prime.
To understand what 5 prime and 3 prime mean, look at the part of DNA’s backbone called the deoxyribose sugar. The carbons are numbered 1 - 5, starting with the carbon that binds the base. Let’s use adenine as an example. The rest are numbered clockwise from this one, so here is the 5 prime carbon. Thymine binds with adenine, and this nitrogenous base is also bound to the 1st carbon on ITS deoxyribose sugar, like this.
Both complementary strands are attached to phosphates, but notice that one of the strands has its 5 prime carbon starting out, and the complementary strand has the 3 prime carbon starting out.
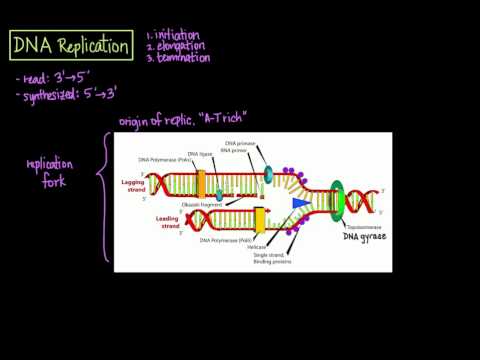
DNA helicase, a wondrous nanomachine, unzips the molecule by breaking the base pair hydrogen bonds. Helicase targets the hydrogen bonds, and prefers bonds between Adenine and thymine since they only have 2 hydrogen bonds, compared with the 3 hydrogen bonds that connect guanine and cytosine. These helicases unzip the DNA molecule in both directions. This point of initial unzipping occurs at the origin of replication.
The helicases continue unzipping the nucleotides to break apart the hydrogen bonds, opening up what is called a replication bubble. So that’s step 1.
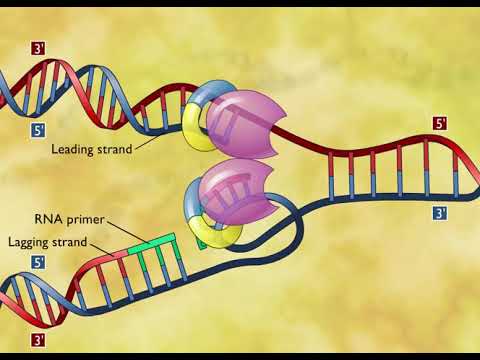
RNA polymerase, our next exquisite nanomachine, adds a few bases, called an RNA primer, to both strands. That is step 2.
Step 3 is when DNA polymerase adds nucleotides in the 5’ to 3’ direction, and it can only add matching nucleotides with nearby ones from the primer to hold onto. That’s why you need to have the RNA primer, since RNA polymerase can add complementary nucleotides all by itself. So DNA polymerase adds these nucleotides continuously in the 5 - 3 direction from the origin of replication, following the ever-moving helicase.
On the complementary strand, DNA polymerase adds nucleotides in the 5 to 3 direction continuously away from origin of replication, following the helicase as it breaks hydrogen bonds. DNA polymerase is also able to edit and correct any mistakes it makes while matching nucleotides from the pool of free nucleotides within the nucleus.
New DNA molecules therefore will end up with one old DNA strand and the brand new strand. This is described as “semi-conservative replication”.
Since DNA can only be added by DNA polymerase in the 5 - 3 direction, more RNA primers need to be added in those parts of the replication bubbles, and while the DNA polymerase still goes in the 5 - 3 direction, the overall movement is still toward the moving helicase, although it lags behind its complementary strand.
We see this same process on the other side of the molecule, as well. Wherever nucleotides cannot be added continuously in the 5 - 3 direction, many more RNA primers are set down, and then DNA polymerase can add nucleotides in the 5 - 3 direction, while still ultimately working its way toward the helicase that is separating more of the molecule. PLay to show another example.
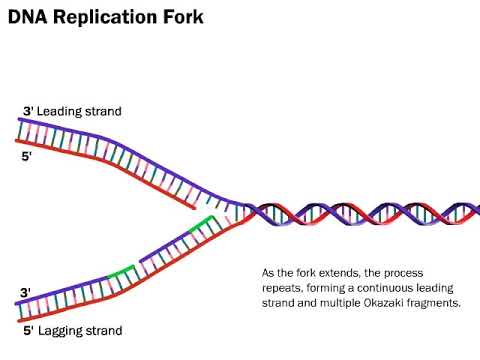
These small fragments formed by the “working backward” replication are called Okazaki Fragments. These are on the lagging strand, so named because it is a little slower to form since it has to work backwards. Compare this with continuous leading strand, which with just one primer, can happily add nucleotides and follow the helicase as it breaks apart hydrogen bonds.
In Step 4, one of the many types of DNA polymerases replaces the RNA primers with DNA, still always working in the 5 to 3 direction. In the last step, DNA ligase, yet another nanomachine, seals all the fragments together.
Another amazing nanomachine called topoisomerase is constantly making little clips in the phosphate sugar backbone to relieve the twisting strain that occurs as helicase unzips DNA.
It then reseals up all these little clips, once the strain on the helix has shifted to another spot, and clips a new place to relieve this torsional strain.
The final product is an identically coded DNA molecule, and these two new molecules remain attached at a DNA sequence called the centromere. These identical DNA molecules are sister chromatids that will separate during mitosis.
z1ozQczaB1M |